 We introduce a revised pipeline called A5-miseq, which replaces several components of the original A5 pipeline with new software modules and produces substantially improved assemblies. The longer reads make it possible to assemble genomes from less data overall, but doing so required major revisions to the data processing algorithms in A5. 
The original A5 could not process reads longer than 150 nt. Since the publication of A5, Illumina’s chemistry has advanced significantly and the MiSeq instruments are now capable of producing reads in excess of 400 nt long, which is 4-fold longer than what was previously possible on a HiSeq 2000. 
The workflow included five steps, and the parameters for each step were optimized on assemblies of Halophilic archaea and tested on E scherichia coli. We previously published A5, a pipeline that automated all the steps to generate bacterial genome assemblies from raw Illumina data ( Tritt et al., 2012). The steps often consist of adapter trimming, quality filtering, error correction, creation of contigs, verification of contigs by mapping reads to the assembly and the creation/verification of scaffolds. Genome assembly involves an entire data processing workflow starting with raw sequence data and ending with scaffolded contigs. Source code and precompiled binaries for Mac OS X 10.6+ and Linux 2.6.15+ are available from Ĭontact: information: Supplementary Data are available at Bioinformatics online. Together, these changes result in substantially improved assemblies that recover a more complete set of reference genes than previous methods.Īvailability: A5-miseq is licensed under the GPL open-source license. Unlike the original A5 pipeline, A5-miseq can use long reads from the Illumina MiSeq, use read pairing information during contig generation and includes several improvements to read trimming. 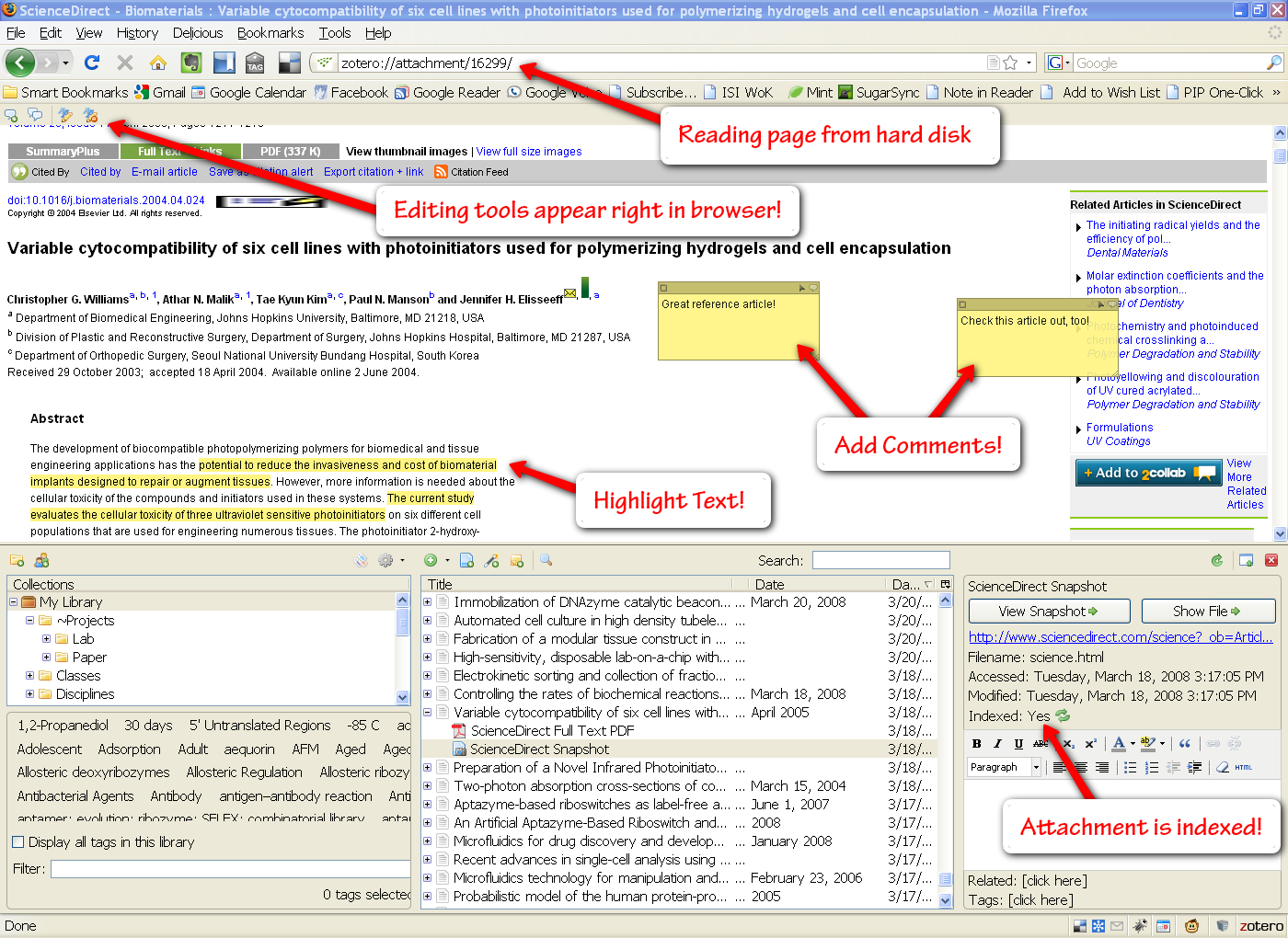
A5-miseq does this by automating the process of adapter trimming, quality filtering, error correction, contig and scaffold generation and detection of misassemblies. Results: A5-miseq can produce high-quality microbial genome assemblies on a laptop computer without any parameter tuning. Few software solutions exist that are capable of automating all steps in the process of de novo genome assembly from Illumina data. Motivation: Open-source bacterial genome assembly remains inaccessible to many biologists because of its complexity. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
